In healthcare, generative AI is transforming how medical professionals analyze data, summarize clinical notes, and generate insights to improve patient outcomes. From automating medical documentation to assisting in diagnostic reasoning, large language models (LLMs) have the potential to augment clinical workflows and accelerate research. However, these innovations also introduce significant privacy, security, and intellectual property challenges.
Healthcare data often contains Protected Health Information (PHI), which is governed by strict regulations and compliance frameworks. At the same time, organizations or researchers who have invested substantial time and compute resources into training medical LLMs must protect their proprietary model architectures, weights, and fine-tuned datasets. Traditional deployment models necessitate mutual trust between the model publisher and the healthcare data provider — trust that sensitive data won’t be leaked, and that the model itself won’t be copied, tampered with, or exfiltrated. The absence of a secure and verifiable trust model between model publishers and consumers remains one of the main barriers to scaling generative AI in regulated medical environments.
To address this concern, both parties need a secure environment to publish and consume models without exposing data or intellectual property. Amazon Web Services (AWS) Nitro Enclaves provide isolated, attested, and cryptographically verified compute environments that help protect sensitive workloads. Model owners can encrypt their LLMs with AWS Key Management Service (AWS KMS) and allow only verified Nitro Enclaves to decrypt and run them, making sure that the model can’t be accessed outside the Nitro Enclave. Healthcare organizations and consumers can use this to process sensitive data within their own AWS environment entirely within the Nitro Enclave, helping keep PHI private and contained. Hardware-based attestation provides proof that the Nitro Enclave is running trusted code, so that both sides can exchange information with confidence.
In this post, we demonstrate how to deploy a publicly available foundational model (FM) using Nitro Enclaves for isolated, more secure compute, AWS KMS for model encryption, Amazon Simple Storage Service (Amazon S3) for storing model artifacts and images, and Amazon Simple Queue Service (Amazon SQS) for securely delivering queries, enabling private, privacy-preserving inferences while helping protect both model intellectual property and sensitive patient data.
This solution outlines how to build a more secure end-to-end pipeline that enables zero trust medical LLM publication and inference with Nitro Enclaves. This post demonstrates a guide for setting up an Amazon Elastic Compute Cloud (Amazon EC2) instance with Nitro Enclaves enabled, downloading and encrypting a publicly available FM to an S3 bucket with an AWS KMS key, sending medical text and image-based queries to an SQS queue for processing, and storing results in an Amazon DynamoDB table.
This project is intended solely for educational and demonstration purposes and isn’t suitable for production or clinical use. Its outputs aren’t validated for clinical accuracy and must not be used for patient care or medical decision-making. Before any real-world deployment, make sure that you implement comprehensive security, privacy, and compliance safeguards. These include health data protection controls, secrets management, and regulatory validation. Furthermore, you must consult the appropriate clinical, legal, and security experts.
For demonstration purposes, this solution is deployed in a single AWS account. Ideally, in production, it would be deployed across separate AWS accounts: one for the model owner and one for the model consumer. The model owner can use cross-account AWS Identity and Access Management (IAM) permissions and encrypted model sharing through AWS KMS to securely provide access to their model without exposing the underlying weights or logic. At the same time, the consumer can run sensitive inferences within their own environment, maintaining strict data privacy and zero trust principles. In a real-world implementation, the model provider should also establish a robust entitlement and licensing framework to manage customer access, enabling fine-grained control over who can invoke the model, track usage, and support license revocation to immediately remove permissions from specific customers when necessary.
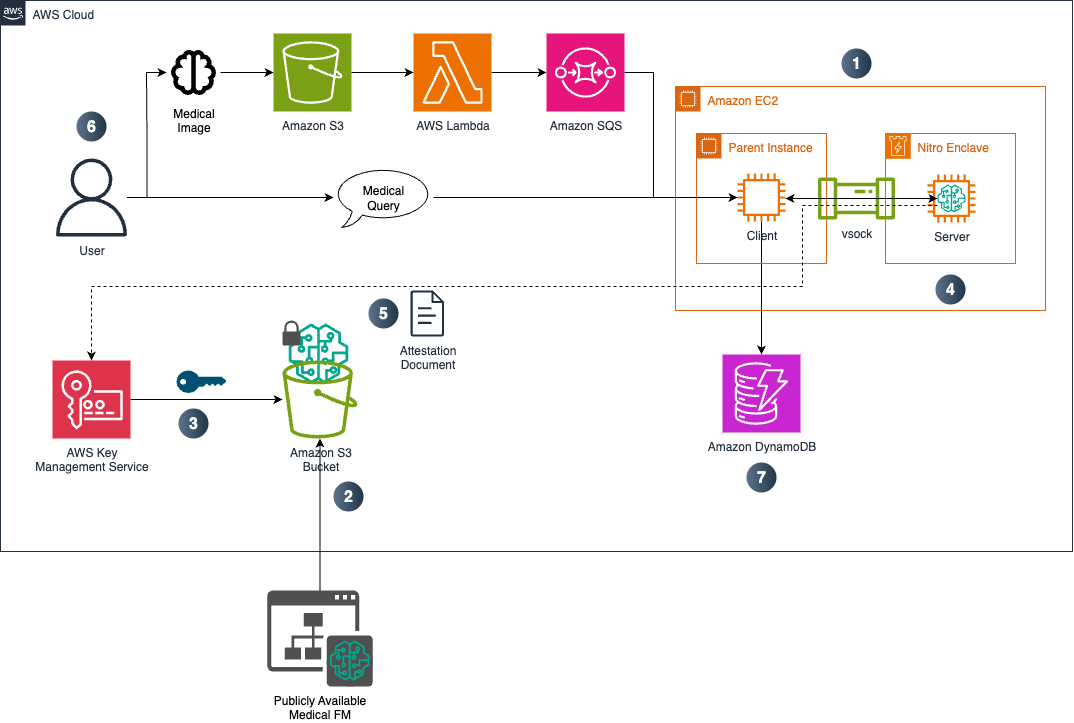
The following diagram shows the solution architecture:
The steps of the solution include:
Google MedGemma is a family of medically-optimized LLMs built on Gemma 3, with 4B and 27B parameter variants supporting both text and multimodal versions for medical image inputs. The 4B model offers efficiency and strong performance for multimodal tasks such as report generation and medical Q&A, while the 27B models excel at more demanding scenarios, such as electronic health record interpretation and complex longitudinal data analysis.
MedGemma models are well-suited for automated radiology report generation, clinical triage and documentation, patient education, medical image pre-interpretation, and medical education systems. The 4B model is ideal for portable or resource-constrained deployments, whereas the 27B multimodal delivers maximal performance.
In this project, MedGemma 4B serves as a reference medical LLM, showing how domain-adapted fine-tuning can enhance a model’s ability to interpret, reason about, and respond to complex medical queries. It also provides a foundation for exploring the safe and effective use of LLMs in healthcare applications, while being securely deployed within a Nitro Enclave. However, you can choose to deploy your own medical FM if needed. This is a deeper overview on the 4B model.
To implement the proposed solution, make sure that you have the following:
The following sections outline how to set up your environment for this solution.
In this solution, you create two S3 buckets: one for the model artifacts and one for the image inputs.
To create the S3 buckets
When images are uploaded to the S3 image bucket, they are sent to an SQS queue for processing in sequential order by the model running in the Nitro Enclave.
To create an SQS queue
For image-based queries, MedGemma 4B expects images encoded in base64 format to be passed in the prompt. To convert the images to this format, a Lambda function is invoked using an Amazon S3 trigger when an image is uploaded to the bucket.
To create a Lambda function
When the queries have been processed by the model for inference, the prompts and responses are logged to a DynamoDB table for auditing and message history purposes.
To create a DynamoDB table
An AWS KMS key is used to envelope-encrypt the model artifacts before they are uploaded to the S3 model bucket. During encryption, the AWS KMS key policy is configured with conditions that restrict decryption to only those Nitro Enclaves presenting a valid attestation document. This attestation includes platform configuration registers (PCR) hashes that represent the measured state of the Nitro Enclave, which covers the signed Nitro Enclave image, runtime, and configuration. When the Nitro Enclave is launched, it generates an attestation document signed by the Amazon EC2 Nitro hypervisor, proving that its PCR values match the expected trusted measurements defined in the AWS KMS key policy. The key is released only if these PCR hashes align and the attestation is verified by AWS KMS, allowing the Nitro Enclave to decrypt and load the model securely in memory.
To create an AWS KMS key
Now that the necessary resources are set up, you can proceed to launch the EC2 instance and create the Nitro Enclave image. For this solution, a c7i.12xlarge instance with a 150 GB EBS volume is provisioned.
To launch an EC2 instance with Nitro Enclaves enabled
With Amazon EC2 loaded with the necessary scripts, you can begin building the Nitro Enclave image. During this process, the Docker container is converted into an Enclave Image File (EIF), which generates cryptographic measurements (PCR hashes) that uniquely identify the code and configuration of the enclave. These measurements are embedded into the AWS KMS key policy, creating a hardware-attested trust boundary that makes sure only this specific, unmodified Nitro Enclave can decrypt and access the model weights.
When the model is decrypted and running on the llama.cpp server within the Nitro Enclave, you can begin to invoke the model with either image or text-based queries. Open a new terminal session in your EC2 instance. You can navigate to the client folder to run the scripts for queries.
For inference on medical images, upload an image to your Amazon S3 image bucket. When it is uploaded, run python3 image_processor.py to pass the image from the SQS queue to the Nitro Enclave for processing. The following are examples of image inputs and model outputs.
Case courtesy of Dr Henry Knipe, Radiopaedia.org, rID: 46289
For inference on text-queries, run python3 direct_query.py "<YOUR_MEDICAL_QUERY>" to invoke the model. The following are examples of text-based inputs and model outputs.
To avoid incurring future charges, delete the resources used in this solution:
You can combine the isolation and attestation capabilities of AWS Nitro Enclaves, the encryption controls of AWS KMS, and the scalability of services such as Amazon S3, Amazon SQS, and Amazon DynamoDB to build a more secure, zero trust pipeline for deploying generative AI models in healthcare. Using Google MedGemma 4B as your reference medical LLM, you can enable privacy-preserving inference where both PHI and model intellectual property remain protected. For more information, consult the following resources: